Preparation of fungi for DNA extraction typically involves growing cultures in liquid culture in Erlenmeyer flasks, Roux bottles or even microfuge tubes (Cenis 1992 Nucl. Acids Res. 20:2380). Growing fungal cultures in liquid may require formulating new media or determining aeration requirements, and there are no rapid means of confirming the identification of the resulting mycelium. Fungi are grown routinely on agar media for identification, but agar complicates DNA extraction.
Solutions of ''reverse agar'' (BASF pluronic polyol F-127), a block polymer of polyoxypropylene and polyoxyethylene, are solid at normal room temperatures, but liquid at 4C. When the compound is used as a replacement for agar in solid media, its unusual properties allow the separation of mycelium and medium by simply placing a mature culture in a refrigerator. The compound has been employed for isolation of heat sensitive antagonistic microorganisms (Gardner and Jones 1984, J. Gen. Microbiol. 130:731-733; Olson and Lange 1989 Opera Bot. 100:197-199), isolation of enzymes associated with basidiome formation in Coprinus (Choi and Ross 1988 Exp. Mycol. 12:80-83), and for isolating mycelium from Neurospora race tubes (Munkres 1990, Fungal Genet. Newsl. 37:26). In this note, the possibility of extracting DNA suitable for PCR amplification from mycelium grown on reverse agar media is documented.
''Reverse Malt Agar'' (RMA) medium was prepared containing 30 BASF pluronic polyol F-127 substituted for agar in the Malt Extract Agar medium of Pitt (1979 The Genus Penicillium, Academic Press). The liquid was poured over the F-127 granules and the resulting suspension left in the refrigerator overnight until dissolved. The solution was then autoclaved, resulting in a thick, sludge like substance, which was again refrigerated, until a homogenous, liquid solution resulted. The liquid was then poured into petri dishes, where it solidified as it warmed to room temperature. The resulting medium is not a true solid, but rather a dense gel.
Cultures of Penicillium spinulosum Thom. (DAOM 216698), Aspergillus japonicus Saito var. japonicus (DAOM 216695), Gliocladium roseum Bainier (Doyle SB-03a, not saved) and Trichoderma harzianum Rifai (DAOM 216501) were grown on RMA for 7 days at 25C in 6 cm petri dishes. Growth rates were slightly slower than on the same medium made with 2 agar, but the resulting colonies produced microscopically typical sporulating structures. For DNA extraction, the petri dishes were placed in the refrigerator for approximately 1 hour until the medium had liquified. Subsequent handling was done on ice. Mycelium was lifted from the medium using an autoclaved pipette tip, placed on the inverted, slanted lid, and allowed to drain for 30-60 seconds. The mycelium was then cut into smaller pieces using a sterile scalpel blade, and transferred into autoclaved 1.5 ml microfuge tubes. The tubes were then spun in a cold microfuge for 5-10 minutes and the excess medium removed. The mycelium was washed twice with 750 uL cold, autoclaved distilled water followed by cold centrifugation, and then used directly for DNA extraction. The DNA miniprep method of Edwards et al. (1991 Nucl. Acids Res. 19:1349), modified by the addition of a cold 70 ethanol wash of the final pellet, was used. The resulting DNA was treated with RNAase A for 1 hour at 37C. PCR amplification of the ITS1-ITS4 region of the ribosomal DNA was performed using the primers and conditions given by White et al. (pp. 282-287 In: PCR Protocols, Innis et al. eds Academic Press). The resulting products were digested for 2 hrs at 37C using HinfI in the buffer supplied with the enzyme.
The DNA yields obtained from mycelium grown on RMA were similar to those obtained from mycelium grown in liquid culture. Washing away excess reverse agar with cold water significantly improved yields, but DNA also was isolated from unwashed mycelium. The resulting DNA performed normally in the ITS PCR amplification and subsequent restriction digests (Figure 1).
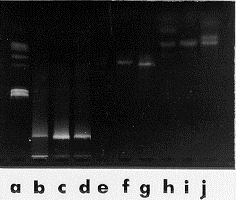
Figure 1. Miniprep DNA (b-d), ITS 1-4 amplification (e-g) and HinfI restriction digests (h-j) from fungi grown on malt extract reverse agar. b, e, h Penicillium spinulosum. c, f, i Aspergillus japonicus var. japonicus. d, g, j Trichoderma harzianum. Lane a is the marker.
Use of reverse agar for cultivation of fungi for DNA extraction may be convenient for certain studies. For fungi that do not produce characteristic structures in liquid broth, reverse agar provides a means of ensuring the correct identity of the mycelium before DNA extraction proceeds. Certain population genetics studies, for example, require the manipulation of a large number of cultures that must be cloned (eg. single-spore isolations) before genetic analysis can proceed. The use of reverse agar could eliminate one round of culture transfers, resulting in significant labour savings.
Reverse agar solutions can be stored for some time at 4C and plates poured when required. Experience has shown that poured plates do not keep indefinitely at room temperature. After 4-6 weeks, the medium no longer liquifies, presumably because of higher polymer concentrations resulting from water evaporation. Also, because the polymer itself is slightly inhibitory to fungi, it does not seem to be suitable for weak nutrient media. Our trials with the Fusarium medium SNA (Nirenberg 1981 Can. J. Bot. 59:1599-1609), for example, resulted in sparse growth that could not be harvested following liquification of the medium.
Acknowledgements: I am grateful to Dr. John Speakman (BASF, Limburghof, Germany) and BASF Performance Chemicals, Parsippany, NJ for providing samples of pluronic polyol F-127.
Note from FGSC: Dr. K.D. Munkres donated a large quantity of pluronic F-127 to the stock center. We will gladly make samples (100-200 g) available available at no cost to interested researchers.
1812年,法国皇帝拿破仑一世从俄罗斯莫斯科撤退时,其大部分军队因饥饿、疾病和寒冷的冬天而损失殆尽。如今,对这撤退途中丧生的30万士兵的部分遗骸的DNA的分析发现,两种未曾预料到的细菌性疾病很可能增加......
1812年夏,法兰西皇帝拿破仑·波拿巴率50万大军入侵俄罗斯帝国。然而到12月时,这支军队仅余零星残部。历史记载将此次“全军覆没”归因于饥寒交迫与斑疹伤寒。但一项新研究表示,从士兵牙齿中提取的DNA,......
美国北卡罗来纳大学研究团队研发出一种名为“DNA花朵”的微型机器人。这种机器人具有独特的自适应环境变化能力,能够像生物体一样,根据周围环境改变形状和行为。“DNA花朵”机器人由DNA与无机材料结合形成......
瑞士苏黎世联邦理工学院科学家在最新一期《自然》杂志上发表论文称,他们开发出一款名为MetaGraph的DNA搜索引擎,能快速、高效地检索公共生物学数据库中的海量信息,为研究生命科学提供了强大的专业工具......
究竟是什么让人脑与众不同?美国加州大学圣迭戈分校研究团队发现了一个名为HAR123的小型DNA片段,这将是解开人类大脑独特性之谜的关键。相关研究成果发表于新一期《科学进展》杂志。最新研究表明,HAR1......
究竟是什么让人脑与众不同?美国加州大学圣迭戈分校研究团队发现了一个名为HAR123的小型DNA片段,这将是解开人类大脑独特性之谜的关键。相关研究成果发表于新一期《科学进展》杂志。最新研究表明,HAR1......
基因组编辑技术作为生命科学领域的一项重要突破,为基础研究和应用开发提供了技术支撑。以CRISPR及其衍生技术为代表的编辑系统通过可编程的向导RNA引导Cas9等核酸酶靶向基因组特定位点,被广泛应用于特......
神经元中基因编辑的插图。图片来源:杰克逊实验室哪怕在五年前,人们也会认为在活体大脑中进行DNA修复是科幻小说中才有的情节。但现在,科学家已能进入大脑、修复突变,并让细胞在整个生命周期中维持住这种修复效......
国际知名学术期刊《自然》北京时间7月2日夜间在线发表一篇基因组学论文称,研究人员从上埃及Nuwayrat地区一个古王国墓葬中提取到一名古埃及个体的全基因组测序数据,这些数据分析可追溯至古埃及第三至第四......
在一项研究中,科学家对埃及一座墓葬中的一名古埃及人进行了全基因组测序。这些数据可追溯至古埃及第三至第四王朝,揭示了其与北非及中东地区,包括美索不达米亚古人群的亲缘关系,为早期埃及人的遗传多样性研究提供......